Ecosystem health shapes viral ecology in peatland soils
Viral communities in UK peatlands
We sampled soils from seven UK peatlands (Fig. 1a and Supplementary Table 1) spanning a gradient of ecosystem health statuses (EHSs): near-natural (undrained/undisturbed reference, hereafter natural), damaged (drained/eroded) and restored (formerly damaged, then rewetted)18. Damaged peatlands were probably drained for decades, although the exact duration is unknown. Restoration age varied by site but was within 10 years of sampling. Compared with natural sites, damaged sites have drier, more oxygenated soils and higher community-wide growth rates18. Restored sites show signs of recovery but remain chemically distinct from natural peatlands18.
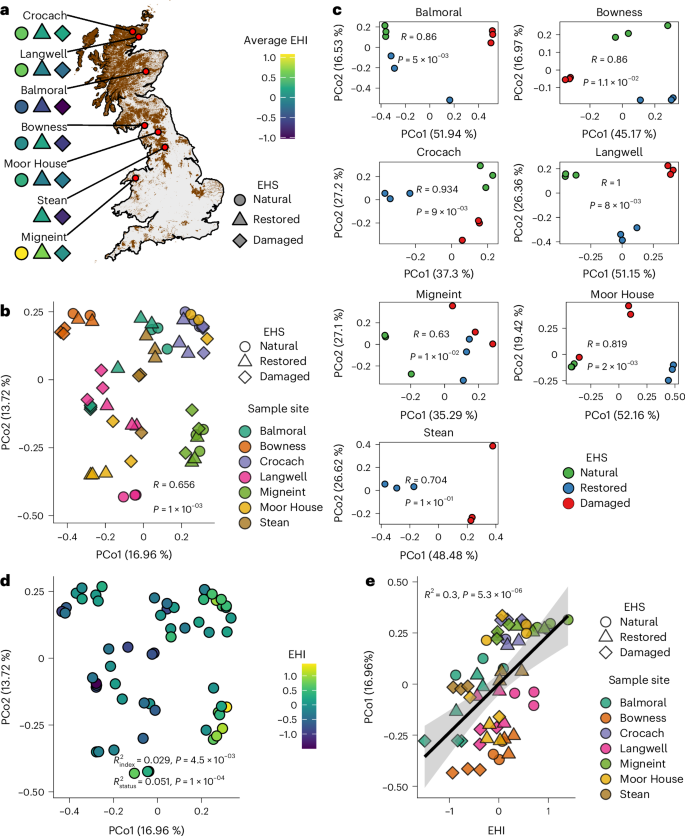
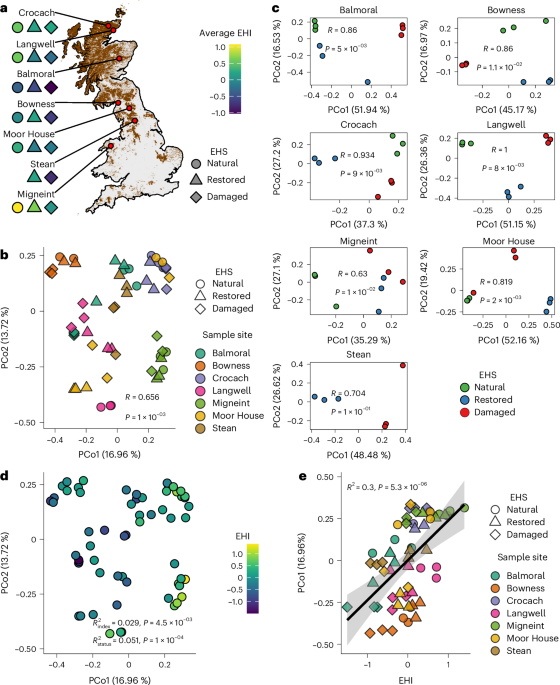
a, Map of the seven peatland sites. Natural, restored and damaged sampling blocks of EHS are coloured by their average EHI. The map shading indicates peatland cover. No natural blocks were sampled at Stean. b, PCoA of viral community Bray–Curtis dissimilarities (n = 60 soil samples). ANOSIM (999 permutations) shows significant separation by site (R = 0.656, P = 1.0 × 10−3, unadjusted). c, Site-specific PCoAs with ANOSIM statistics for separation of communities by EHS (999 permutations; exact R and P values shown on plots). d, The PCoA from b coloured by EHI. PerMANOVA (two-sided, 999 permutations) reports the marginal R2 for variance in community dissimilarity explained by EHI (R2index = 0.029, P = 4.5 × 10−3 BH-adjusted) and EHS (R2status = 0.051, P = 1.0 × 10−4, BH-adjusted). e, Linear regression of PCoA axis 1 against EHI, where the black line represents the fitted regression mean and the shaded band indicates the 95% confidence interval around the fitted line. Regression statistics from a linear model of the two axes are provided (R2 = 0.30, P = 5.28 × 10−6).
Source data
We sequenced community DNA of soil samples and co-assembled metagenomes by combining triplicate sequence read libraries from each sampling site and EHS, yielding 22 assemblies (one per site × EHS combination) of high quality (Supplementary Tables 1 and 2). We identified 3,177 viral scaffolds across all sites and EHS from metagenome co-assemblies, which were binned into 2,281 viral metagenome-assembled genomes (vMAGs) (Supplementary Table 3). These genomes were dereplicated and clustered into 1,548 virus species-level clusters which were analysed downstream using virus species-level representative genomes.
Environmental differences between EHSs
A principal components analysis (PCA) across sites revealed that EHS generally reflected the composition of environmental parameters (Extended Data Fig. 1a). PCA loadings indicated that total carbon (0.72), pH (0.63) and oxygen concentration (0.61) were the strongest parameters, with moisture (0.57), total nitrogen (0.51) and conductivity (0.38) being also important, supporting EHS’s role in structuring peatland soils. Site-specific effects were strong, as shown by separate PCAs within sites, where the relative influence of environmental variables differed (Extended Data Fig. 1b). To capture the complex variation across sites and EHSs, an ecosystem health index (EHI) was previously calculated for each sample18 (Fig. 1a), incorporating peat chemistry, oxygen, moisture and vegetation. This index provides a holistic, continuous measure of peatland ecosystem health that effectively reflects variation across all samples.
While EHS grouped samples within sites, the degree of separation varied, suggesting that local environmental conditions matter. This is consistent with the fact that these sites span a climatic gradient and vary in their level of degradation and restoration18. Nevertheless, soils from damaged peatlands were less waterlogged, more oxygenated and more acidic compared with the natural contrast where soils from restored peatlands demonstrated signs of mitigation. Overall, these results indicate that while site-level differences are prominent, EHS captures meaningful environmental variation across peatland sites. Likewise, changes in EHI offer a useful means to track relative improvements in ecosystem health when comparing areas of varying EHSs within a given peatland.
Geography and ecosystem health structure peatland virus communities
To explore the drivers of viral community composition, we performed principal coordinate analysis (PCoA) and found that geography was the primary structuring factor (Fig. 1b). Although the viral communities of some samples from different sites were similar in composition, samples in the PCoA were mostly grouped by their geographic origin (R = 0.656, P = 0.001, analysis of similarities (ANOSIM)). The influence of EHS on community structure became more apparent when analysing sites separately. Within sites, we observed a strong separation of samples by EHS (R > 0.5, P < 0.05, ANOSIM; Fig. 1c). The exception was Stean, where lack of natural reference samples probably reduced power. Alongside overall EHS grouping, we mapped the previously calculated EHI for each sample18 to our PCoA (Fig. 1d) and found that EHI also significantly impacted viral community structure (R2 = 0.029, P = 4.5 × 10−3, PerMANOVA; Supplementary Table 4) independently of EHS (R2 = 0.051, P = 1 × 10−4). EHI was also strongly positively correlated with virus community PCo1 (Fig. 1e), providing further evidence that ecosystem health is a significant factor that drives viral community structure. Host community composition (Extended Data Fig. 2) also significantly impacted viral community structure, but this did not overshadow the independent effects of sample site and ecosystem health (Supplementary Table 4 and Supplementary Results).
Peatland soils contain a mix of endemic and shared viral populations
Considering the strong effects of geography and ecosystem health on structuring environmental variation across sites, we examined the degree of endemism among our identified virus species. Most viral genomes were detected in soil metagenomes from multiple sample sites (76% of species representatives, Fig. 2a). However, 54% of virus species were endemic to individual EHS (found exclusively in one of natural, damaged or restored soils across all sites) compared with 46% that were shared (Fig. 2b). We also assessed whether the viral genomes identified were largely novel or instead represented in published soil virus databases. We gathered a comprehensive collection of genomes from the three largest and most recent soil virus databases17,19,20. We found that more viruses from this study formed genus-level genome clusters with viruses from other databases than other viruses from this study (Fig. 2c). Thus, many viral genomes clustered with known soil viral genomes from other ecosystems, indicating that not all are unique to peatlands at the genus level. These results suggest that soil viruses in UK peatlands share a core of virus lineages with other soils, alongside a substantial fraction of locally endemic viruses.
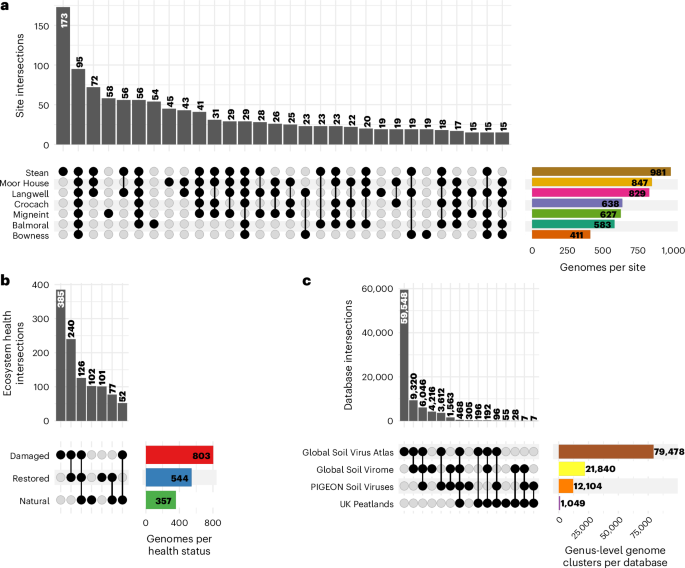
a,b, Detection of species-representative virus genomes across soils from different sample sites (a) and soil with different EHSs (b). For simplicity, intersections with <15 genomes are omitted in a and b. c, Intersections of databases represented in genus-level clusters of the viruses here and other genomes from three soil virus databases17,19,20 (n = 729,998 genomes). Bars represent clusters with genomes originating from each database in the intersection. Numbers above or inside bars indicate the total number of genome clusters with genomes originating from each database in the intersection.
Source data
Viruses are differentially abundant across EHSs
Having established that ecosystem health significantly shapes viral communities, we next identified viruses that were differentially abundant across EHSs. Using DESeq2 (ref. 21), we created ecosystem health ‘trend groups’ for a qualitative analysis of functions (Supplementary Results and Extended Data Fig. 3). Host genomes were also differentially abundant across EHSs and were clustered into trend groups (EHS group; Supplementary Results and Extended Data Fig. 4). For detailed information on the distribution and clustering of viral species-representative genomes, see Supplementary Results. Across all sites, there was a greater proportion of damaged-enriched viruses (37%) than restored-enriched viruses (33%) and natural-enriched viruses (29%) (Fig. 3a). This contrasted with trends for hosts, indicating that damaged peatlands host a greater share of enriched viruses among differentially abundant groups. In summary, the differential abundance of viral species across EHSs shows that environmental health strongly influences viral population sizes, which vary strongly between natural, restored and damaged peatland soils.
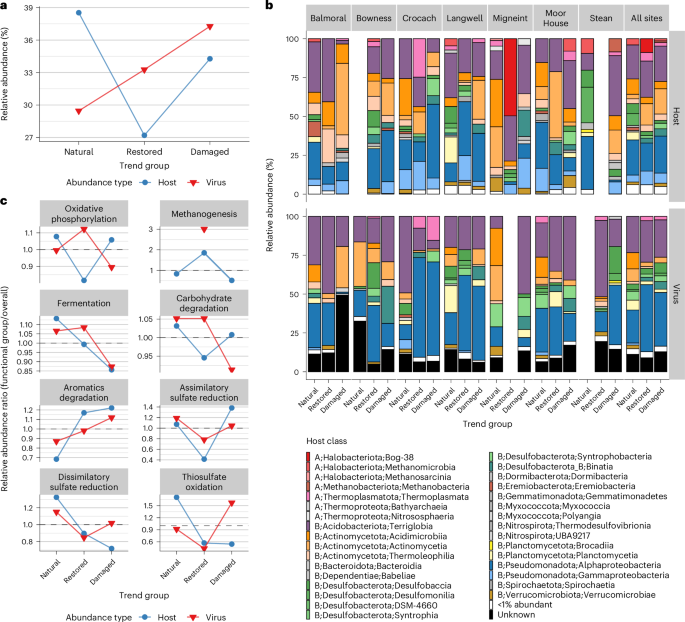
a, The relative abundance of virus (n = 1,448 genomes) and host (n = 411 genomes) genomes differentially abundant across EHS ‘trend groups’. b, The relative abundance of host genomes (n = 411 host genomes, top row) and virus genomes (n = 1,351 virus genomes, bottom row), categorized by host class. Only differentially abundant host and viral genomes are included. Host classes are labelled as follows: Domain (A = Archaea, B = Bacteria); Phylum; Class. Individual classes that were <1% in relative abundance were grouped into the same ‘<1% abundant’ category. c, Abundance ratios of differentially abundant hosts encoding eight key metabolic functions relevant to peat soils (n = 407 host genomes) and their predicted viruses (n = 550 virus genomes).
Source data
The abundance of viruses across EHSs is discordant with dominant host taxa
We examined the relative abundance of differentially abundant bacterial and archaeal MAGs and their predicted viruses at the host class level within each EHS (Fig. 3b). Relative abundances of viruses and their hosts varied substantially across sites. However, when examining all sites together, viruses infecting hosts in the phyla Actinomycetota, Desulfobacterota and Planctomycetota showed a marked decrease in abundance from the natural to the restored trend groups. This was met with an increase in viruses infecting Pseudomonadota hosts, particularly Alphaproteobacteria. Viruses of Alphaproteobacteria and Desulfobaccia also increased in abundance from natural to damaged trend groups. These viral abundance shifts did not mirror host changes. For example, while the relative abundance of Pseudomonadota viruses surged from natural to restored groups, the abundance of Pseudomonadota hosts remained stable. Similarly, Desulfobaccia hosts made up only 1.8% of damaged-enriched hosts, yet Desulfobaccia viruses represented 6.6% of damaged-enriched viruses. These findings show that viral and host dynamics across EHSs are discordant, suggesting that viral responses to environmental changes may depend on factors beyond host availability.
Viral and host dynamics across key biogeochemical functions in peatlands
Given that peatlands at different EHSs are chemically distinct, we explored whether viruses infecting microbes with key biogeochemical functions changed across EHSs, and whether these changes reflected overall viral and host abundance trends. We calculated the relative abundance of viruses infecting hosts with eight metabolic functions (Supplementary Table 5) within each EHS group, normalized by overall viral abundance within that same trend group (Fig. 3c). Similar calculations were done for predicted hosts. Below, we focus on notable trends, but comprehensive results for all eight metabolic functions are provided in Supplementary Results.
Across the eight metabolic functions, several key trends stood out. For oxidative phosphorylation, viral abundance (n = 451 virus genomes) increased from natural (1.00) to restored soils (1.12, +11%), but decreased again from restored to damaged (0.89, −25%). Host abundance (n = 365 host genomes), in contrast, decreased from natural to restored (0.82, −32%). For fermentation, viral abundance (n = 491 virus genomes) remained stable between natural (1.07) and restored soils (1.08, +1.7%) but showed a decrease in damaged soils (0.87, −22%), paralleling a similar decrease in host abundance (−32%, n = 477 host genomes). Carbohydrate degradation showed minor changes, with viral abundance (n = 542 virus genomes) being stable between natural and restored soils (1.05) but decreasing slightly in damaged soils (0.91, −15%), alongside minor fluctuations in host abundance (n = 590 host genomes). For assimilatory and dissimilatory sulfate reduction, there were decreases in viral abundance (−52% and −36%, n = 20 and n = 105 virus genomes) from natural to restored soils, accompanied by an even greater decrease for their hosts (−159% and −48%, n = 34 and n = 68 host genomes). From restored to damaged soils, viruses infecting hosts with these functions increased by 25% (assimilatory) and 17% (dissimilatory). Although assimilatory sulfate-reducing hosts showed a major increase of 70% from restored to damaged soils, dissimilatory sulfate-reducing hosts declined by 25% over the same transition, contrasting with the pattern observed for their viruses. A similar pattern was observed for thiosulfate oxidation (n = 33 virus genomes, n = 16 host genomes). It is important to note that these percentage changes reflect descriptive trends based on aggregated ratios and were not subjected to null hypothesis testing (see Methods). These patterns suggest that while virus and host dynamics often align, the enrichment of hosts with specific metabolic functions, such as oxidative phosphorylation and sulfur cycling, can sometimes diverge from the enrichment of viruses that infect them across different EHSs.
Viral proteins are functionally distinct across EHSs
To assess viral functional differences across EHSs at the protein level, we clustered protein-coding viral genes from all sites and examined their distribution across EHSs (Fig. 4a). The three largest groups were protein clusters unique to individual EHS, suggesting that soils of each EHS harbour viruses encoding proteins with distinct functions. This degree of adaptation is notably greater than what we previously observed at the genome level (Fig. 2b). These patterns indicate that common pools of viral genomes exist across EHSs, but their functional potential is locally adapted to their specific environmental conditions. Despite these functional distinctions, the distribution of functional categories (based on PHROG22) across the intersections remains consistent (Fig. 4a). Therefore, while viruses are specialized at the protein level, they perform similar high-level functions across all EHSs. Overall, our results demonstrate that viral protein functions are finely tuned to their environments, even when broader categories are conserved.
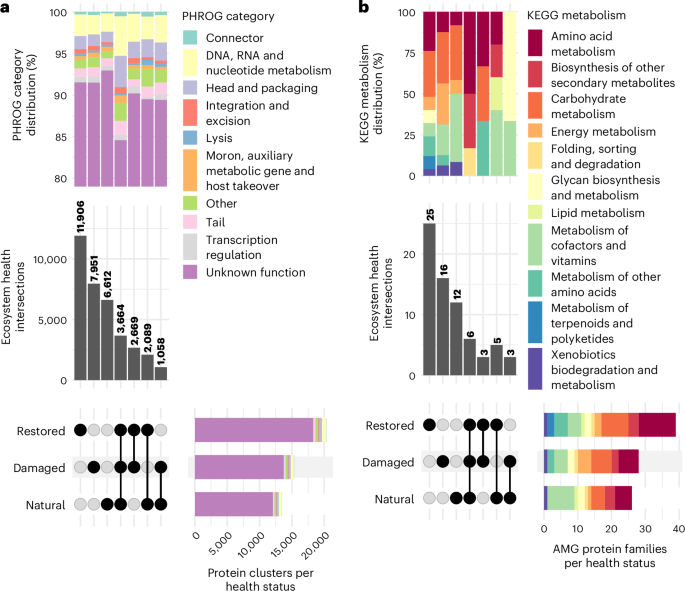
a, UpSet plot showing amino acid identity-based clustering of all viral protein-coding genes (n = 77,662 genes). Intersections represent protein clusters with viral proteins from multiple EHSs, while non-intersecting groups represent proteins unique to a single EHS. The distribution of PHROG22 functional categories across these intersections is shown in the stacked bar plot at the top. b, UpSet plot of unique KEGG KOfams24 (n = 59 families) among viral AMGs (n = 100 genes) across EHSs. Intersections indicate KOfams shared across different EHSs, while non-intersecting groups highlight KOfams unique to a single EHS. The stacked bar plot at the top illustrates the distribution of KEGG metabolism categories associated with these viral KOfams across the intersections.
Source data
We also focused on viral AMGs23 and their distribution across EHSs (Fig. 4b). AMGs are host-derived proteins with metabolic functions that provide viruses with evolutionary and fitness benefits. Similar to the all-protein results, the largest intersections correspond to KEGG24 protein families unique to individual EHS, reinforcing the idea that the metabolic functions encoded by these viral genomes are distinct across different environmental conditions. Likewise, the distribution of high-level KEGG metabolism categories across the major intersections remained largely similar, with categories such as ‘Carbohydrate metabolism,’ ‘Metabolism of cofactors and vitamins’ and ‘Amino acid metabolism’ being well represented. Yet, there was a small increase in the proportion of energy metabolism genes in the damaged-only samples compared with the natural-only and restored-only samples, with predicted functions involved in sulfur metabolism (K20034 3-(methylthio)propionyl-CoA ligase), methane metabolism (K16370 6-phosphofructokinase 2 and K15229 methylamine dehydrogenase heavy chain) and oxidative phosphorylation (K02107 V/A-type H+/Na+-transporting ATPase subunit G/H). This subtle shift may indicate functional adaptation, with viruses in damaged, oxygenated soils potentially playing a more active role in processes linked to electron transport in their hosts for their selfish benefit25,26. Altogether, viral proteins and AMGs are not distinct at high-level functions across EHSs, yet they are locally adapted to specific restoration contexts.
Virus–host infection dynamics change with EHS
Viruses are dependent on their hosts to replicate, but their modes and rates of replication vary27,28. To this end, we next investigated virus–host infection dynamics using genome abundances of our bioinformatically predicted virus–host pairs. Linear regressions between total virus abundance and total host abundance across EHSs reveal complex interactions that vary by phylum (Fig. 5a). Notably, while the slopes of these regressions change within each phylum depending on EHS, all slopes are consistently less than 1. For example, in Acidobacteriota, the slopes are 0.567 (R2 = 0.84, BH-adjusted P = 2.67 × 10−7, n = 18 soil samples) in natural soils, 0.812 (R2 = 0.55, BH-adjusted P = 3.81 × 10−4, n = 19 soil samples) in restored soils and 0.719 (R2 = 0.84, BH-adjusted P = 3.19 × 10−6, n = 15 soil samples) in damaged soils, indicating that host genomes are generally more abundant than their associated viral genomes across all EHSs at the phylum level. This pattern suggests chronic or non-lytic modes of infection at high host densities, known as ‘piggyback-the-winner’ dynamics28,29, where viruses coexist with their hosts through non-lethal replication strategies, such as lysogeny, involving integration into the host genome. This pattern was also observed for viruses and hosts of other dominant phyla, but the strengths of these relationships were susceptible to changes in EHS (see Supplementary Results).
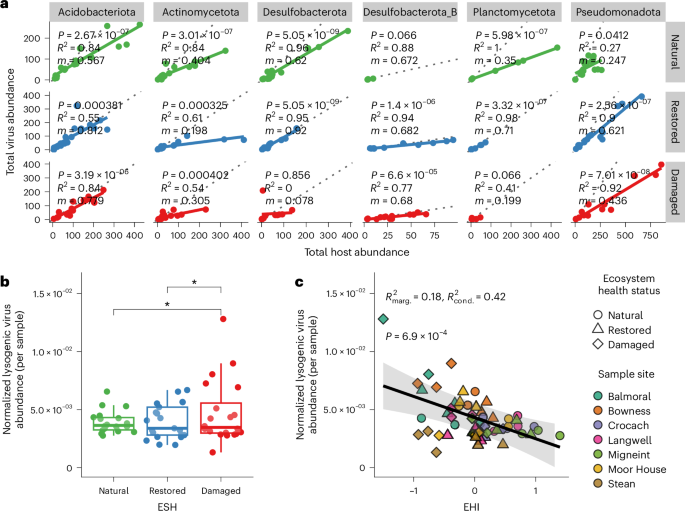
a, Linear regressions of virus and host abundances by EHS. For clarity, only the six host phyla with the highest number of observations are shown. Linear model statistics (two-sided) for each phylum × EHS combination are provided, where m is the slope of the best-fit line (shown as a solid line representing the fitted regression mean) and P gives BH-adjusted P values for the significance of each regression slope. Dotted lines represent a hypothetical slope of m = 1. b, Average trimmed mean genome coverage of lysogenic viruses per sample (normalized by the total trimmed mean coverage of all virus genomes; n = 60 soil samples) across all sites, grouped by EHS. Significant pairwise contrasts among EHSs are shown (estimated marginal means, two-sided, *P ≤ 0.01, BH-adjusted), determined from a linear mixed-effects model with sample site as a random intercept. Boxplots: centre line, median; box limits, upper and lower quartiles; whiskers, 1.5× interquartile range; points, individual data points. c, Linear mixed-effects model predicting normalized lysogenic virus abundance per sample in b (n = 60 soil samples) from EHI, with site as a random intercept. The black line shows the marginal fitted values (population-level mean predictions) from the linear mixed-effects model, and the shaded band represents the corresponding 95% confidence intervals. The marginal (R2marg. = 0.18) and conditional (R2cond. = 0.42) R2 of the model fit are shown, and P = 6.9 × 10−4 (unadjusted) reflects the result of a Type II ANOVA assessing the significance of EHI as a fixed effect in the model.
Source data
Lineage-specific shifts in lysogeny and induction across EHSs
While ‘piggyback-the-winner’ dynamics prevailed in UK peatland soils, patterns of temperate (hereafter lysogenic) and actively replicating viral abundance across EHSs highlighted significant shifts in virus–host interactions. We identified 297 lysogenic viruses in total, 13% of all identified viruses, and analysed their abundances in each sample. Patterns of lysogenic virus abundance varied across sites, with no significant differences in their raw mean abundances when aggregating all sites (Extended Data Fig. 5). However, when normalizing lysogenic virus abundance by the total virus population in each sample (Fig. 5b), we found that the proportion of lysogenic viruses was significantly lower in natural and restored soils compared with damaged soils (estimated marginal means, BH-adjusted P = 0.0300 and P = 0.0398, respectively). This suggests that lysogenic viruses contributed more substantially to the overall viral community in damaged soils. Furthermore, when modelling normalized lysogenic virus abundance as a function of the EHI while accounting for site-level variation (Fig. 5c), we observed a significant negative relationship (marginal R2 = 0.18, conditional R2 = 0.42, χ2 = 11.52, BH-adjusted P = 6.9 × 10−4, Type II ANOVA, n = 60). This indicates that the relative abundance of lysogenic viruses increases with peatland degradation. Together, these findings suggest an increase in the replication of lysogenic viruses as peatlands shift from natural to damaged states.
We aimed to identify actively replicating viruses in our samples by calculating virus-to-host abundance ratios (also known as virus:microbe ratio, or VMR) (Extended Data Fig. 6). We considered a virus to be ‘active’ if the virus:host ratio exceeded 10. Using this threshold, we identified 51 active viruses across 46 samples. This represented 10% of all viruses with host predictions and non-zero virus and host abundances. Of the 51 active viruses, 27 (53%) were also predicted to be lysogenic, accounting for 9.1% of all predicted lysogenic viruses. Thus, these active lysogenic viruses probably underwent recent induction at the time of soil sampling. Among them, 26% were active in natural soils, 41% in restored soils and 67% in damaged soils (13 lysogenic viruses were active in more than one sample, explaining why the total exceeds 100%). We also found that EHS had a significant effect on virus:host ratios, but the effects varied by the host family (see Supplementary Results). In summary, these results support our observation that both overall viral genome abundance and the proportion of lysogenic virus genomes increase in damaged soils, with a subset of these viruses probably undergoing greater induction and replication compared with those in natural and restored peatlands.
link







